This R package (Hahsler, Piekenbrock, and Doran 2019) provides a fast C++ (re)implementation of several density-based algorithms with a focus on the DBSCAN family for clustering spatial data. The package includes:
Clustering
Outlier Detection
Fast Nearest-Neighbor Search (using kd-trees)
The implementations use the kd-tree data structure (from library ANN)
for faster k-nearest neighbor search, and are typically faster than the
native R implementations (e.g., dbscan in package fpc), or
the implementations in WEKA, ELKI and Python’s scikit-learn.
The following R packages use dbscan: AFM, bioregion, CLONETv2, ClustAssess,
cordillera,
CPC, crosshap, daltoolbox, DDoutlier, diceR, dobin, doc2vec, dPCP, EHRtemporalVariability,
eventstream,
evprof, FCPS, fdacluster, FORTLS, funtimes, FuzzyDBScan,
karyotapR, ksharp, LOMAR, maotai, metaCluster,
mlr3cluster,
MOSS, oclust, openSkies, opticskxi, OTclust, pagoda2, parameters, ParBayesianOptimization,
performance,
rMultiNet, seriation, sfdep, sfnetworks, sharp, shipunov, smotefamily,
snap, spdep, spNetwork, squat, ssMRCD, stream, supc, synr, tidySEM, weird
To cite package ‘dbscan’ in publications use:
Hahsler M, Piekenbrock M, Doran D (2019). “dbscan: Fast Density-Based Clustering with R.” Journal of Statistical Software, 91(1), 1-30. doi:10.18637/jss.v091.i01 https://doi.org/10.18637/jss.v091.i01.
@Article{,
title = {{dbscan}: Fast Density-Based Clustering with {R}},
author = {Michael Hahsler and Matthew Piekenbrock and Derek Doran},
journal = {Journal of Statistical Software},
year = {2019},
volume = {91},
number = {1},
pages = {1--30},
doi = {10.18637/jss.v091.i01},
}Stable CRAN version: Install from within R with
install.packages("dbscan")Current development version: Install from r-universe.
install.packages("dbscan",
repos = c("https://mhahsler.r-universe.dev",
"https://cloud.r-project.org/"))Load the package and use the numeric variables in the iris dataset
library("dbscan")
data("iris")
x <- as.matrix(iris[, 1:4])DBSCAN
db <- dbscan(x, eps = 0.42, minPts = 5)
db## DBSCAN clustering for 150 objects.
## Parameters: eps = 0.42, minPts = 5
## Using euclidean distances and borderpoints = TRUE
## The clustering contains 3 cluster(s) and 29 noise points.
##
## 0 1 2 3
## 29 48 37 36
##
## Available fields: cluster, eps, minPts, metric, borderPointsVisualize the resulting clustering (noise points are shown in black).
pairs(x, col = db$cluster + 1L)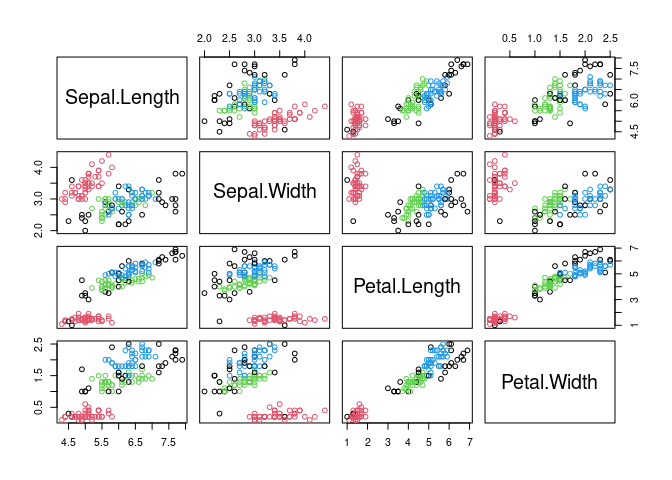
OPTICS
opt <- optics(x, eps = 1, minPts = 4)
opt## OPTICS ordering/clustering for 150 objects.
## Parameters: minPts = 4, eps = 1, eps_cl = NA, xi = NA
## Available fields: order, reachdist, coredist, predecessor, minPts, eps,
## eps_cl, xiExtract DBSCAN-like clustering from OPTICS and create a reachability plot (extracted DBSCAN clusters at eps_cl=.4 are colored)
opt <- extractDBSCAN(opt, eps_cl = 0.4)
plot(opt)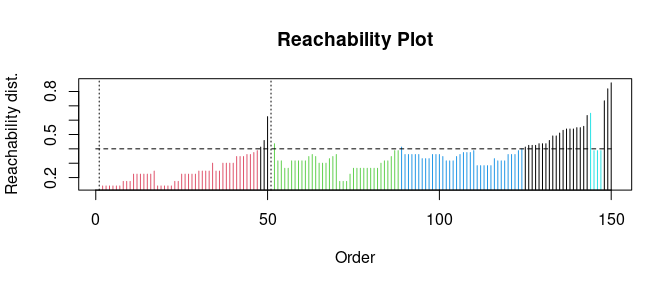
HDBSCAN
hdb <- hdbscan(x, minPts = 4)
hdb## HDBSCAN clustering for 150 objects.
## Parameters: minPts = 4
## The clustering contains 2 cluster(s) and 0 noise points.
##
## 1 2
## 100 50
##
## Available fields: cluster, minPts, coredist, cluster_scores,
## membership_prob, outlier_scores, hcVisualize the hierarchical clustering as a simplified tree. HDBSCAN finds 2 stable clusters.
plot(hdb, show_flat = TRUE)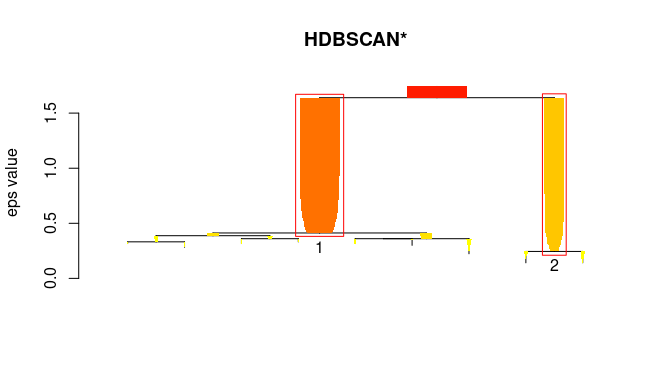
dbscan provides for all clustering algorithms
tidy(), augment(), and glance()
so they can be easily used with tidyverse, ggplot2 and tidymodels.
library(tidyverse)
db <- x %>%
dbscan(eps = 0.42, minPts = 5)Get cluster statistics as a tibble
tidy(db)## # A tibble: 4 × 3
## cluster size noise
## <fct> <int> <lgl>
## 1 0 29 TRUE
## 2 1 48 FALSE
## 3 2 37 FALSE
## 4 3 36 FALSEVisualize the clustering with ggplot2 (use an x for noise points)
augment(db, x) %>%
ggplot(aes(x = Petal.Length, y = Petal.Width)) + geom_point(aes(color = .cluster,
shape = noise)) + scale_shape_manual(values = c(19, 4))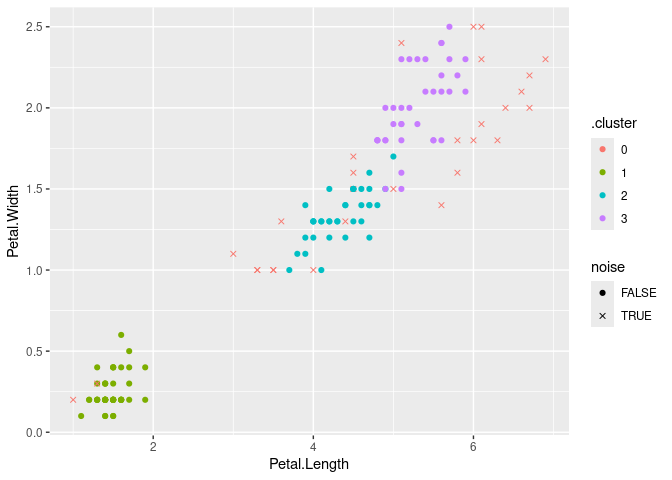
R, the R package dbscan, and the Python package
rpy2 need to be installed.
import pandas as pd
import numpy as np
### prepare data
iris = pd.read_csv('https://archive.ics.uci.edu/ml/machine-learning-databases/iris/iris.data',
header = None,
names = ['SepalLength', 'SepalWidth', 'PetalLength', 'PetalWidth', 'Species'])
iris_numeric = iris[['SepalLength', 'SepalWidth', 'PetalLength', 'PetalWidth']]
# get R dbscan package
from rpy2.robjects import packages
dbscan = packages.importr('dbscan')
# enable automatic conversion of pandas dataframes to R dataframes
from rpy2.robjects import pandas2ri
pandas2ri.activate()
db = dbscan.dbscan(iris_numeric, eps = 0.5, MinPts = 5)
print(db)## DBSCAN clustering for 150 objects.
## Parameters: eps = 0.5, minPts = 5
## Using euclidean distances and borderpoints = TRUE
## The clustering contains 2 cluster(s) and 17 noise points.
##
## 0 1 2
## 17 49 84
##
## Available fields: cluster, eps, minPts, dist, borderPoints# get the cluster assignment vector
labels = np.array(db.rx('cluster'))
labels## array([[1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1,
## 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 0, 1, 1,
## 1, 1, 1, 1, 1, 1, 2, 2, 2, 2, 2, 2, 2, 0, 2, 2, 0, 2, 2, 2, 2, 2,
## 2, 2, 0, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 0,
## 2, 2, 2, 2, 2, 0, 2, 2, 2, 2, 0, 2, 2, 2, 2, 2, 2, 0, 0, 2, 0, 0,
## 2, 2, 2, 2, 2, 2, 2, 0, 0, 2, 2, 2, 0, 2, 2, 2, 2, 2, 2, 2, 2, 0,
## 2, 2, 0, 0, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2]],
## dtype=int32)The dbscan package is licensed under the GNU General Public License (GPL) Version 3. The OPTICSXi R implementation was directly ported from the ELKI framework’s Java implementation (GNU AGPLv3), with permission by the original author, Erich Schubert.